Wheeler Lab Folks
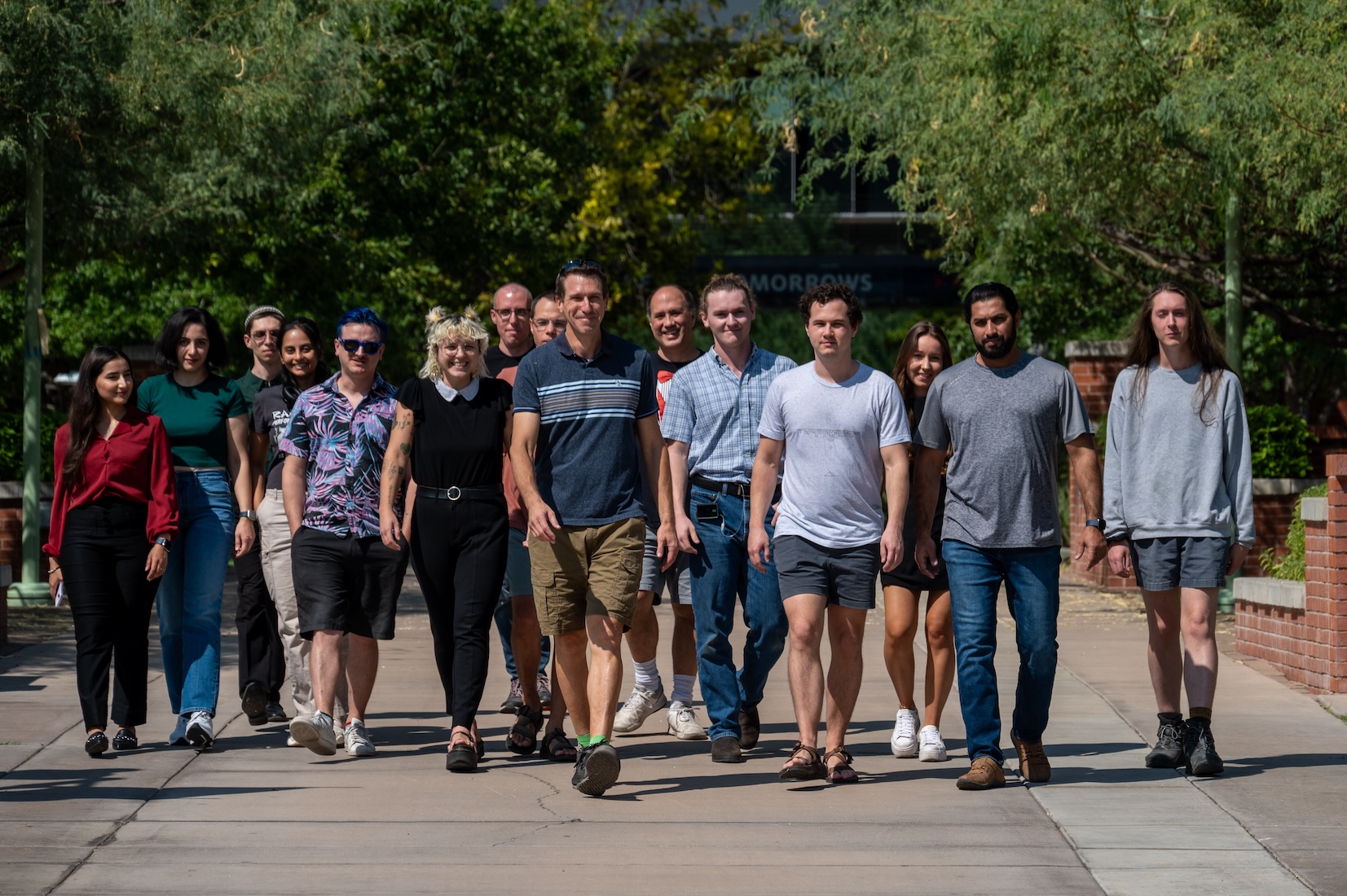
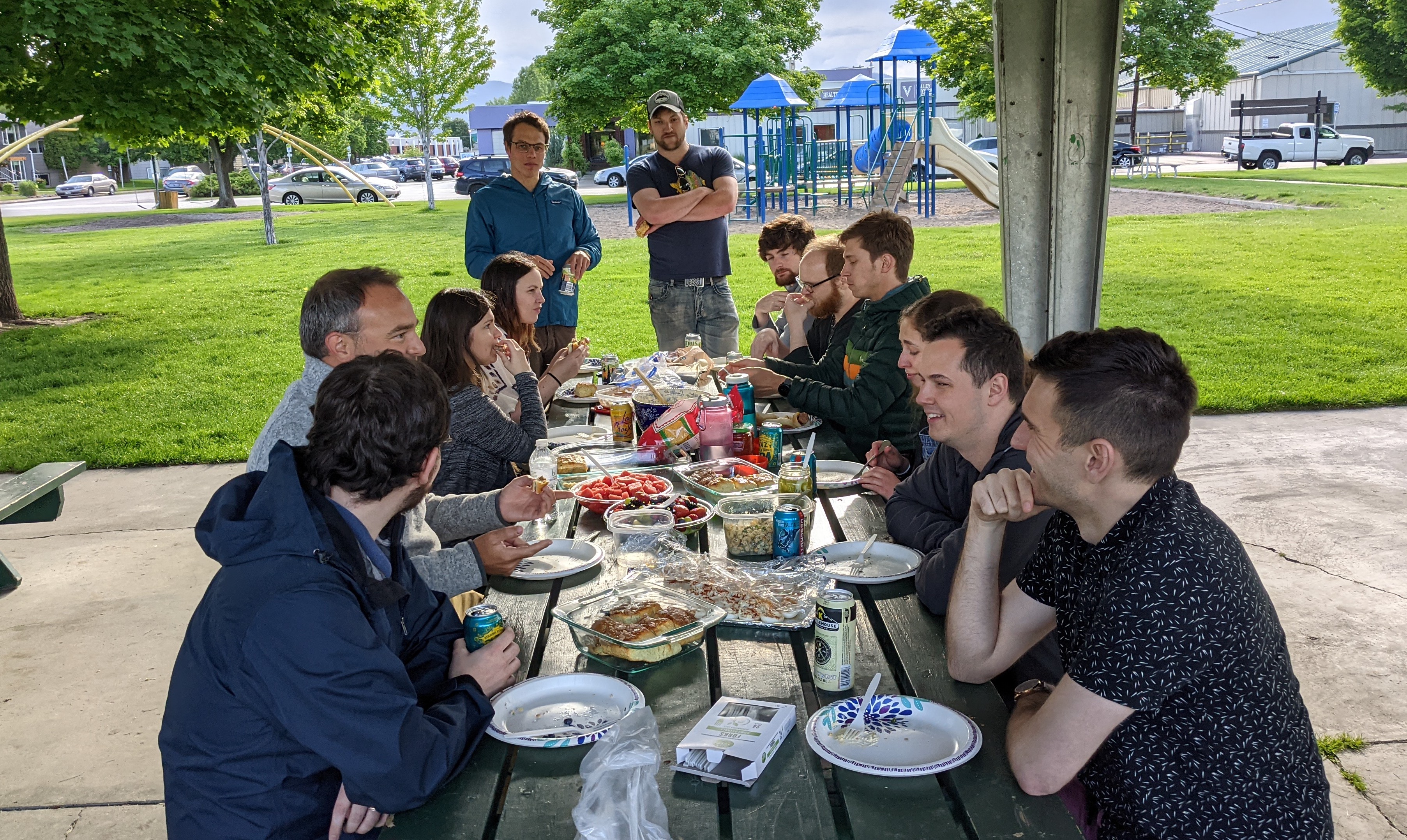




Travis Wheeler

Chief Suggestion Officer and Manuscript Bottleneck
Email | Personal Page | Curriculum Vitae | Google
Scholar profile | 
University of Arizona
Department
of Pharmacy Practice & Science
Affiliate:
Computer
Science, School of Information
Applied Mathematics, Statistics
and Data Science
Genetics, Bio5
Editorial Board Member - Bioinformatics
Advances, Mobile
DNA
Advisory Board Member - Montana IDeA Network of Biomedical
Research Excellence
Research: Computational genomics, Computational drug discovery, Machine Learning
Research staff
Clément Goubert

Research Professor – Email
| Personal Page
Transposable elements, evolution, splicing, and all things genomics
Jack Roddy

Senior Software Scientist – Email | Personal Page
Jack of all trades, master of some
(including sequence
annotation and visualization)
Ken Youens-Clark

Senior Software Scientist – Email | Personal Page
Software and web/cloud services
Genevieve Krause

Software Scientist – Email | Personal Page
Sequence annotation with profile HMMs; frameshift model for protein-coding DNA.
George Glidden

Software Scientist – Email | Personal Page
Peptide and modification identification in Mass Spectrometry; animal tracking.
Terrill Yuhas

Senior Data Architect – Email
| Personal Page
(Joint
with Bonnie LaFleur)
Building data systems for precision medicine research
Isaac Robinson

Software Scientist – Email | Personal Page
Algorithms
for sequence annotation; ML methods for tracking and behavior
classification.
Postdocs
(None at the moment; maybe you?)
PhD students
Anna Marbut

PhD candidate, Computer Science (DIS) (UMontana) (2019-) – Email | Personal Page
Research: Language representation (ML and NLP)
BA, Linguistics, Reed College
MA, Business Analytics, University
of Montana
Jeremiah Gaiser

PhD student, School of Information (2023-) – Email | Personal Page
Previously:
Undergraduate researcher, Biochemistry, (2017-2020); Biochem PhD student
in Wheeler lab at UMontana (2020-2023) (transfered to UofA)
Research: Protein-drug biding affinity prediction
BS, Computational Biochemistry, University of Montana
Neha Sontakke

PhD student, Computer Science (2023-) – Email | Personal Page
Research: Algorithms and statistical models for metabolic network gap filling
BS, Computer Science, Pune Institute of Computer Technology
Dylan Wichman

PhD student, Computer Science (2026-) – Email | Personal Page
Research: Deep Learning for drug discovery
BS, Computer Science, LSU
Masters students
(None at the moment; maybe you?)
Undergraduate students
(None at the moment; maybe you?)
Lab Alumni
Regular Collaborators
Arian Smit (Institute for Systems Biology)
Robert Hubley (Institute for Systems Biology)
Travis Hughes (University of Montana)
Jason McDermott (Pacific Northwest National Laboratory)
Nathan Insel (Wilfrid Laurier University)